
Principal Investigator
In our brain, axons transmit information over long distances, and are susceptible to degenerate during aging and in pathological conditions. We study how axons are preserved during life and how mitochondrial integrity power axonal function.
We are interested in understanding how axons maintain their function during a lifetime. Human axons can reach the length of 1 m, and contain up to 99% of the cytoplasm. Axonal function and survival depend on efficient trafficking mechanisms to deliver proteins, lipids, and cellular organelles, including mitochondria, to their distal regions. Neuronal survival critically depends on the integrity and functionality of mitochondria. We aim to understand which cellular pathways are implicated in axonal degeneration. Moreover, we investigate how the quality and the function of mitochondria is regulated and maintained in axons during life.
The Rugarli group’s research is also dedicated to understanding how the viability of axons in healthy people is maintained over the course of many decades. One approach to tackle this question is to understand the function of genes that contribute to the pathogenesis of genetic diseases characterized by axonal degeneration (for instance heredity spastic paraplegia, HSP). The researchers’ ultimate vision is to identify common signal cascades triggered in the axons that would allow for interventions. A second approach is to elucidate processes that maintain the functionality of distal axonal organelles, such as the endoplasmic reticulum and the mitochondria in axons. Decoding the process of axonal degeneration is not only important for a better understanding of HSP. The results could also translate to other aging-associated neurodegenerative diseases such as Parkinson’s or Alzheimer’s.
Axons are like long and narrow roads connecting a main factory to outposts located far away. Avoiding traffic accidents and maintaining the efficiency of the outposts is essential during our lifetime. Understanding how this is achieved is at the core of our research.
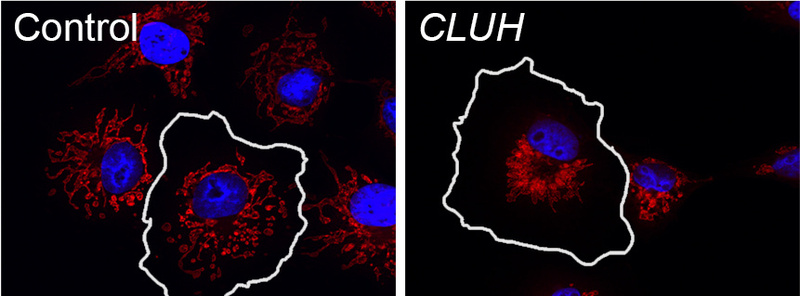
Our lab has contributed to the understanding of the molecular and cell biological function of several HSP genes, and pinpointed the pathogenic role of defective microtubule-severing, lipid droplet metabolism, and mitochondrial quality control in the degenerative process. In addition, the group is studying a post-translational mechanism to regulate mitochondrial biogenesis, which involves the cytosolic RNA-binding protein CLUH. CLUH binds nuclear-encoded mRNAs for mitochondrial proteins and control their stability and translation. The ultimate goal is to achieve a deeper understanding of the processes that maintain mitochondrial quality in neurons and theor long axonal processes.
Our current aims include:

Principal Investigator